Welcome to Xcelris Labs Ltd. GitHub (by Bioinformatics division)
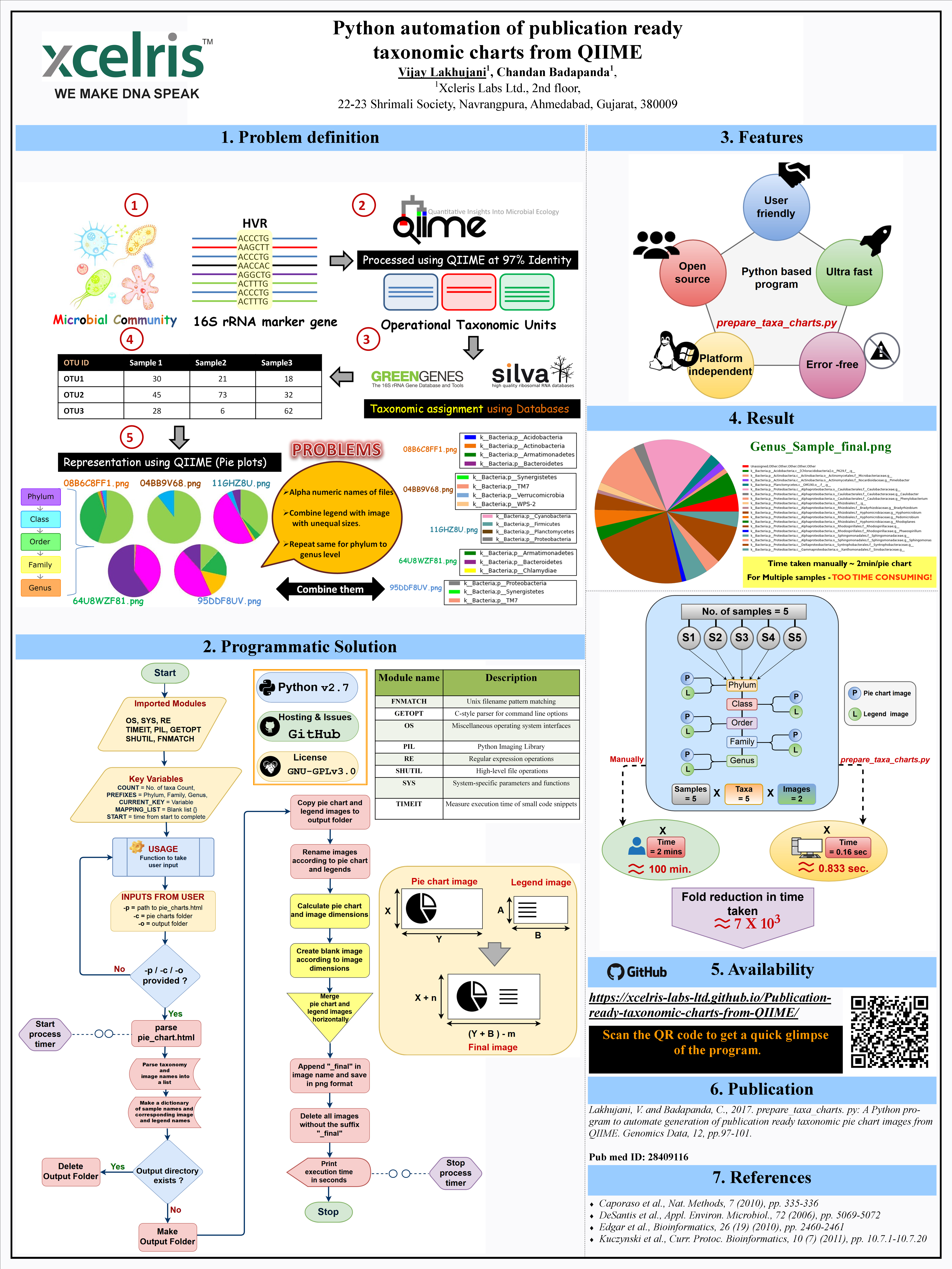
prepare_taxa_charts.py: Ultrafast program to generate publication ready taxonomic pie chart images from QIIME

-
This script parses the pie_chart.html file (output from
plot_taxa_summary.py) -
Maps the pie chart and legend image names to corresponding samples.
-
Copies the files to user defined directory
-
Rename the files according to sample ids
-
Merges the pie charts and legends
Usage
python prepare_taxa_charts.py -p pie_charts.html -c /charts_folder -o user_defined_output_folder
List of Python modules used
| S. No. | Module |
|---|---|
| 1. | fnmatch |
| 2. | os |
| 3. | getopt |
| 4. | PIL |
| 5. | re |
| 6. | shutil |
| 7. | sys |
| 8. | timeit |
Important Note
Prerequisites for prepare_taxa_charts.py:
-
Python v2.7. The program will not work with Python v3. Python v2.7 was chosen as it is stable.
-
Legend files should be generated in
pngformat (default is pdf). This could be achieved by running the scriptplot_taxa_summary.pywith the option-t/--type_of_fileaspng. Given below is an example:
plot_taxa_summary.py -i phylum.txt -l phylum -c pie -o phylum_charts/ -t png
** Note: High resolution images can be generated using the -d, --dpi option for the script plot_taxa_summary.py. See below example (setting the dpi to 100 produce good quality images):
plot_taxa_summary.py -i phylum.txt -l phylum -c pie -o phylum_charts/ -t png -d 100
License
This script is released under GNU GENERAL PUBLIC LICENSE, v3, 29 June 2007
Development and Maintenance
Developed by - Vijay Lakhujani:

Project Scientist, Bioinformatics
NGS division, Xcelris Labs Ltd
vijay.lakhujani@xcelrislabs.com
Let’s connect!
Maintained by @Xcelris-Labs-Ltd on GitHub.
Publication
Citing
Please cite: Lakhujani, V. and Badapanda, C., 2017. prepare_taxa_charts. py: A Python program to automate generation of publication ready taxonomic pie chart images from QIIME. Genomics Data, 12, pp.97-101.
Official entry for poster presentation
The work was formally presented as scientific poster in Accelerating biology-2018 symposium held during Jan, 9-11, 2018 at C-DAC, Pune, India